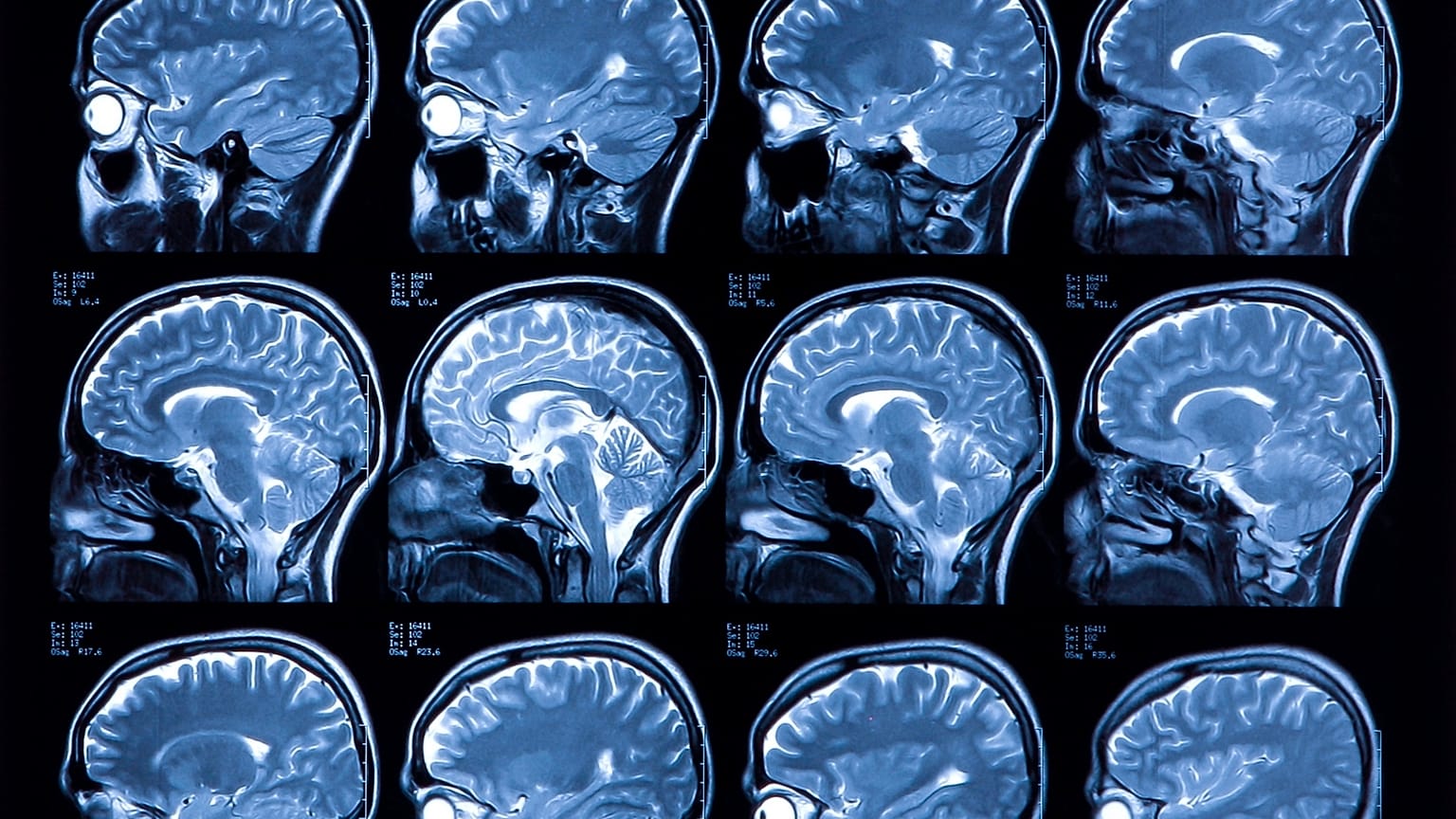
A groundbreaking deep learning study can label 100,000 MRI scans for cancer and neuro patients in minutes rather than years.
Magnetic Resonance Imaging, or MRI scanning, is crucial to diagnosing a plethora of medical conditions; from cancer to neurological disease.
 ADVERTISEMENT
ADVERTISEMENT
 ADVERTISEMENT
ADVERTISEMENT
But accurately labelling each image taken during an examination can be time-consuming, having an impact on how quickly medical staff can correctly identify an illness.
Now researchers at King’s College in London have made a breakthrough in automating the process which means up to 100,000 MRI brain examinations can be labelled in a matter of 30 minutes rather than the years it would take to complete manually.
The research team at the university’s School of Biomedical Engineering and Imaging Sciences had to teach machine learning image recognition systems through so-called "deep learning," which involves multiple algorithms processed through several neural networks.
This was done by training the systems to obtain the labels from radiology reports and correctly assign them to their corresponding MRI examinations.
King's College said the study is the first to enable researchers to label complex MRI datasets at scale, details of which were published in European Radiology.
According to the researchers, deep learning typically requires processing tens of thousands of labelled images in order to achieve the best possible image recognition performance.
Large datasets equal bottlenecks
But the process represents a bottleneck in the development of deep learning systems for complex image datasets.
"By overcoming this bottleneck, we have massively facilitated future deep learning image recognition tasks and this will almost certainly accelerate the arrival into the clinic of automated brain MRI readers," Dr Tom Booth, a senior author on the study, said.
"The potential for patient benefit through, ultimately, timely diagnosis, is enormous".
He added that the models were evaluated for using both unseen labels as well as images.
"While this might seem obvious, this has been challenging to do in medical imaging because it requires an enormous team of expert radiologists. Fortunately, our team is a perfect synthesis of clinicians and scientists," Booth said.
Dr David Wood, the study’s lead author, said the research builds on recent breakthroughs in natural language processing, which had used huge collections of unlabeled text including all of the English language Wikipedia and (open-access research repository) PubMed Central abstracts and full-text articles.
"In the spirit of open-access science, we have also made our code and models available to other researchers to ensure that as many people benefit from this work as possible," he added.